The clinical utility of genomic data has yet to be fully realised. We have been an early adopter of machine learning methods for genomic prediction of phenotypes and have led the design of software tools and the comparison of various competing methodologies to select the right approach for the right disease. As a proof-of-concept, we have shown that a genomic risk score for celiac disease exhibits maximum prediction compared to other related approaches and demonstrates clinically useful information which can be used to remodel the celiac diagnostic pathway (Abraham et al. 2014).
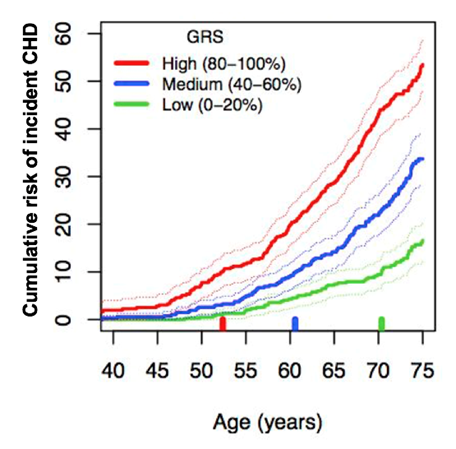 For coronary heart disease, we've published a genomic risk score of 49,000 SNPs which can stratify the top 20 per cent of men who are at high lifetime risk, leading to disease 12–18 years earlier than men at the bottom 20 per cent of risk (Abraham et al. 2016). These high-risk individuals could be candidates for early intervention. Towards this end, the genomic risk score for coronary heart disease is currently being trialled internationally in studies to motivate individuals to change their lifestyle and traditional cardiovascular risk factors. Genomic prediction has profound ramifications for celiac disease, coronary heart disease and other complex diseases and, if properly developed, would likely have immensely positive implications for individual risk stratification and lifestyle intervention to halt pathogenesis in its earliest stages.
For coronary heart disease, we've published a genomic risk score of 49,000 SNPs which can stratify the top 20 per cent of men who are at high lifetime risk, leading to disease 12–18 years earlier than men at the bottom 20 per cent of risk (Abraham et al. 2016). These high-risk individuals could be candidates for early intervention. Towards this end, the genomic risk score for coronary heart disease is currently being trialled internationally in studies to motivate individuals to change their lifestyle and traditional cardiovascular risk factors. Genomic prediction has profound ramifications for celiac disease, coronary heart disease and other complex diseases and, if properly developed, would likely have immensely positive implications for individual risk stratification and lifestyle intervention to halt pathogenesis in its earliest stages.
Relevant reading
Genomic prediction of coronary heart disease
European Heart Journal 2016 ehw450
Accurate and robust genomic prediction of celiac disease using statistical learning
PLoS Genetics 2014 10(2): e1004137
Performance and robustness of penalized and unpenalized methods for genetic prediction of complex human disease
Genetic Epidemiology 2013 37(2):184–95